the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
Whole-soil warming decreases abundance and modifies the community structure of microorganisms in the subsoil but not in surface soil
Nicholas O. E. Ofiti
Jennifer L. Soong
Emily F. Solly
Margaret S. Torn
Arnaud Huguet
Guido L. B. Wiesenberg
Michael W. I. Schmidt
The microbial community composition in subsoils remains understudied, and it is largely unknown whether subsoil microorganisms show a similar response to global warming as microorganisms at the soil surface do. Since microorganisms are the key drivers of soil organic carbon decomposition, this knowledge gap causes uncertainty in the predictions of future carbon cycling in the subsoil carbon pool (> 50 % of the soil organic carbon stocks are below 30 cm soil depth).
In the Blodgett Forest field warming experiment (California, USA) we investigated how +4 ∘C warming in the whole-soil profile to 100 cm soil depth for 4.5 years has affected the abundance and community structure of microorganisms. We used proxies for bulk microbial biomass carbon (MBC) and functional microbial groups based on lipid biomarkers, such as phospholipid fatty acids (PLFAs) and branched glycerol dialkyl glycerol tetraethers (brGDGTs). With depth, the microbial biomass decreased and the community composition changed. Our results show that the concentration of PLFAs decreased with warming in the subsoil (below 30 cm) by 28 % but was not affected in the topsoil. Phospholipid fatty acid concentrations changed in concert with soil organic carbon. The microbial community response to warming was depth dependent. The relative abundance of Actinobacteria increased in warmed subsoil, and Gram+ bacteria in subsoils adapted their cell membrane structure to warming-induced stress, as indicated by the ratio of anteiso to iso branched PLFAs. Our results show for the first time that subsoil microorganisms can be more affected by warming compared to topsoil microorganisms. These microbial responses could be explained by the observed decrease in subsoil organic carbon concentrations in the warmed plots. A decrease in microbial abundance in warmed subsoils might reduce the magnitude of the respiration response over time. The shift in the subsoil microbial community towards more Actinobacteria might disproportionately enhance the degradation of previously stable subsoil carbon, as this group is able to metabolize complex carbon sources.
- Article
(3181 KB) - Full-text XML
- BibTeX
- EndNote
Soil temperatures are predicted to increase at all soil depths in near synchrony with air temperatures under global climate change (Soong et al., 2020a). Global model simulations predict a likely increase in air temperatures between 2.6 and 4.8 ∘C by 2100 under Representative Concentration Pathway 8.5 (IPCC, 2013). It remains uncertain how this temperature increase will affect the soil organic carbon (SOC) pool. Soil organic carbon contains 2 to 3 times the amount of carbon present in the atmosphere (SOC ∼1500–2400 Pg C; IPCC, 2013) and subsoils (below 30 cm soil depth) store around 50 % of the total SOC (Jobbagy and Jackson, 2000). Microorganisms are key players in the SOC cycle, but in subsoils there still exist large uncertainties on how the microbial community will respond to warming (Jansson and Hofmockel, 2020). Environmental constraints for microorganisms are different in subsoils and topsoils, including plant inputs, SOC availability and stabilization processes, soil moisture, or spatial heterogeneity (Rumpel and Kögel-Knabner, 2011). These constraints in subsoils generally lead to less microbial biomass and a microbial community which is adapted to a lower carbon availability (Brewer et al., 2019; Fierer et al., 2003b). Thus, topsoil and subsoil microbial communities may respond differently to warming, with unknown effects on subsoil SOC stocks.
So far only a few field studies have been conducted to test the temperature sensitivity of subsoil microbial activity in situ (Hicks Pries et al., 2017; Nottingham et al., 2020), whereas there have been many incubation and mesocosm studies (e.g., Bai et al., 2019; Fierer et al., 2003a; Lin et al., 2018). The latter indicate that subsoil respiration is equally or more sensitive to warming compared to topsoils (Bai et al., 2019; Fierer et al., 2003a; Lin et al., 2018), consistent with in situ observations of field warming experiments (Hicks Pries et al., 2017; Nottingham et al., 2020). The underlying microbial processes in subsoils are largely unexplored. To the best of our knowledge, only Zhang et al. (2015) investigated how in situ warming over several years affected subsoil microbial biomass and community structure in a grassland experiment. They reported that warming increased microbial biomass throughout the whole-soil profile and observed a depth-specific change in the microbial community structure (Zhang et al., 2015). In topsoils, microbial abundance tends to respond to warming differently, depending on the ecosystem (Chen et al., 2015). Consequently, data from non-grassland ecosystems are needed to see whether the responses reported by Zhang et al. (2015) can be confirmed at other sites.
Processes controlling the microbial response to warming are still subject to intense scientific debate (Walker et al., 2018). An interplay of acclimation and substrate depletion has been proposed to control the microbial response to warming (Melillo et al., 2017; Pold et al., 2017; Walker et al., 2018). Acclimation to warming encompasses physiological adaptations of individual microorganisms and shifts in the microbial community composition (Bradford et al., 2008). Substrate depletion, on the other hand, is a consequence of increased microbial activity with warming, which causes carbon limitation and, in turn, negatively affects microbial processes (Kirschbaum, 2004). But how these processes control the warming response of microorganisms in subsoils remains largely unexplored until now.
We can assess microbial biomass and functional community composition using several parameters. Microbial biomass carbon (MBC) quantified by chloroform fumigation extraction is a proxy for overall microbial biomass (Vance et al., 1987) but does not yield any information on the microbial community composition. Phospholipid fatty acids (PLFAs) are microbial membrane lipids that can be used to quantify microbial biomass and also to describe the microbial community composition at a low taxonomic resolution (Willers et al., 2015). Furthermore, ratios among specific PLFAs have been used as indicators of environmental stress in soil (Wixon and Balser, 2013). For example, the ratio of anteiso to iso branched PLFAs is used as an indicator of a physiological adaptation of the microbial membrane of Gram positive (Gram+) bacteria to stress caused, among others, by changes in temperature (Wixon and Balser, 2013). Adjusting the abundance of these compounds in their membrane helps microorganisms to control the fluidity of the membrane to maintain their activity under changing environmental conditions (Hall et al., 2010). We chose PLFA analysis over techniques with a higher resolution because of the precision between replicates and the high statistical power in differentiating experimental treatment effects (Ramsey et al., 2006). Core lipids of branched glycerol dialkyl glycerol tetraethers (brGDGTs) are an independent and complementary molecular marker that largely represents microbial necromass (Gocke et al., 2017). brGDGTs generally have a longer turnover time compared to the faster cycling PLFAs (Kindler et al., 2009; Weijers et al., 2010), giving a more time-integrated signal of microbial abundance. Our first observations from the study site indicate that there might be less microbial necromass in warmed plots but relatively more microorganism-derived fatty acids compared to plant-derived fatty acids, especially in the subsoil below 50 cm (Ofiti et al., 2021). Thus, we used brGDGTs as a complementary proxy to explore potential changes in microbial necromass. This combination of molecular proxies for microbial abundance has not been explored so far in the context of manipulative soil warming experiments.
In this study, we make use of a soil warming experiment that warms the soils by +4 ∘C above ambient temperature down to 1 m depth at the University of California Blodgett Experimental Forest (Sierra Nevada, CA). We quantified how a temperature increase throughout the soil profile influenced the microbial biomass and community structure, using microbial biomarkers.
Our research was guided by the following three hypotheses
First, we hypothesized that warming would lead to more microbial biomass in top- and subsoils. Warming increased soil respiration at all depths at the study site (Hicks Pries et al., 2017; Soong et al., 2021), which might be partially reflected in more microbial biomass. This would be in line with findings that, in topsoil, microbial biomass generally increases over the first years of soil warming, despite microbial biomass in forest soils being less responsive compared to other ecosystems (Chen et al., 2015). Second, we hypothesized that microbial necromass, as assessed by brGDGTs, will decrease despite the expectation that the microbial biomass will increase. This hypothesis is based on the observation that microorganism-derived fatty acids were less abundant in warmed plots at our study site and that soil organic carbon was more decomposed (Ofiti et al., 2021). Thus, warming might accelerate microbial necromass turnover. Third, we hypothesized that warming-induced changes in the microbial community composition will be depth dependent. As previously reported for soil profiles, we expect that the microbial community composition of top- and subsoils is also different at our site because the subsoil microbial community is adapted to carbon scarcity and other environmental constraints (Brewer et al., 2019; Fierer et al., 2003b). A meta-analysis proposed that the abundance of Actinobacteria increases with warming (Chen et al., 2015). Actinobacteria are relatively more abundant in subsoils compared to topsoils (Fierer et al., 2003b), and an increase in this group would affect the subsoil microbial community more than the topsoil community.
2.1 Experimental setup and sampling
A whole-soil warming experiment was initiated in 2013 at the University of California Blodgett Experimental Forest, Sierra Nevada, CA (120∘39′40′′ W, 38∘54′43′′ N), to study how soil warming affects biogeochemical cycles in a natural environment (Hicks Pries et al., 2017). The site is located in a Mediterranean climate, with a mean annual precipitation of 1660 mm and a mean annual air temperature of 12.5 ∘C. The soil was classified as Alfisol of granitic origin and has a developed O horizon. The experiment is located in a coniferous forest with ponderosa pine (Pinus ponderosa), sugar pine (Pinus lambertiana), incense cedar (Calocedrus decurrens), white fir (Abies concolor), and Douglas fir (Pseudotsuga menziesii) as dominant tree species. The experiment consists of three blocks, each containing a pair of circular control and warmed plots 3 m in diameter. All plots are equipped with vertical steel pipes which contain resistance heater cables and sand in warmed plots and sand only in control plots. Additionally, two rings of heating cables were buried at 5 cm to account for surface heat loss. With this setup, temperature was maintained at +4 ∘C from 0.2–1 m and +2.6 ∘C above 0.2 m in the warmed plots compared to the control plots (Hicks Pries et al., 2017; Soong et al., 2021). The temperature magnitude was slightly lower above 0.2 m due to surface heat loss. Diurnal and seasonal temperature variations were considered depth specifically (Hicks Pries et al., 2017; Soong et al., 2021). Soil samples were collected in April 2018, after 4.5 years of warming, with a 4.78 cm diameter manual soil corer. Organic horizons were removed prior to sampling. Mineral soil samples were recovered sequentially in 10 cm increments down to 90 cm depth and stored at 4 ∘C for transportation to the laboratory. In the laboratory, samples were freeze-dried, sieved to 2 mm, and fine roots were removed before conducting subsequent analyses. Carbon concentrations were measured on an elemental analyzer coupled to an isotope ratio mass spectrometer (EA-IRMS; FLASH 2000-HT Plus, linked by ConFlo IV to DELTA V Plus IRMS; Thermo Fisher Scientific) on a ground subsample. The pH was measured on a dried subsample in 0.01 M CaCl2. A second core was sampled from each plot, using the same methods, and stored at 4 ∘C until chloroform fumigation extraction was conducted during the week after sampling.
2.2 Microbial parameters
2.2.1 Microbial biomass carbon
Microbial biomass carbon (MBC) was analyzed using chloroform fumigation extraction, which is a method that measures the carbon released upon lysis of microbial cells (Vance et al., 1987). The extraction was conducted within 1 week after sampling on field moist soil and stored at 4 ∘C until processing. Due to a limitation of sample material, the analysis was only conducted on the 0–10, 10–20, 30–40, 50–60, and 80–90 cm depth increments using 10 g of the field-moist soil. Microbial biomass carbon was calculated as the difference in total organic carbon between subsamples fumigated with chloroform for 5 d prior to extraction with 0.05 M K2SO4 and subsamples directly extracted with 0.05 M K2SO4. Dissolved organic carbon was measured on a Shimadzu TOC-L (Shimadzu Corporation) and calculated on a soil dry mass basis. A correction factor of 0.45 was used to account for carbon losses inherent to the method and to make data comparable to previously published results (Vance et al., 1987). The difference in MBC between warmed and control plots was calculated as the difference between the warmed and control treatments normalized to the control plot within each of the three replicate paired blocks.
2.2.2 Phospholipid fatty acids
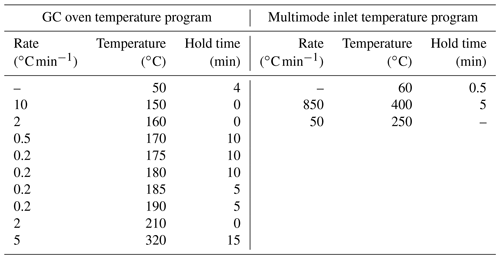
Analysis of PLFAs was performed following the method by Frostegård et al. (1991), with some modifications (Zosso and Wiesenberg, 2021). Briefly, we increased the amount of freeze-dried soil sample with soil depth to account for the decreasing microbial biomass (from 3 g in the top soil to 12 g in deepest depth intervals). Using a solution of () of chloroform : methanol : citric acid buffer, the extraction was repeated four times and the extract combined in a separation funnel. After the addition of citric acid buffer and chloroform, the phases separated overnight, and the organic phase was released thereafter. We added chloroform three more times and combined the organic phases. The extract was then separated in neutral-, glyco-, and phospholipid fractions using a column with activated silica gel, and the phospholipid fraction was methylated using a boron trifluoride-methanol solution (Wiesenberg and Gocke, 2017). Prior to methylation, deuterated eicosanoic acid (D39C20; Cambridge Isotope Laboratories, Inc.) was added as an internal standard for quantification to nanomoles per gram of soil (hereafter, nmol g−1). Quantification was performed using a gas chromatograph (GC; 7890 B; Agilent Technologies, Inc.) equipped with a multimode inlet (MMI) and a flame ionization detector. External standard series of 24 fatty acid standards and sample measurements on a GC (6890 N; Agilent Technologies, Inc.) coupled to a mass spectrometer (5973 N; Agilent Technologies, Inc.), followed by comparison to the National Institute of Standards and Technology (NIST) and Wiley mass spectra library, supported peak identification. Both GCs were equipped with J&W DB-5MS columns (Agilent Technologies, Inc.) with the following dimensions: 50 m length × 0.2 mm inner diameter × 0.32 µm film thickness. The GC oven and MMI temperature programs are given in the Appendix (Table B1).
For total abundance of PLFAs, saturated fatty acids (C14:0, C15:0, C16:0, C17:0, and C18:0) were included. They were not included in calculations of the relative abundances of microbial groups (mole percentage; hereafter mol%) because they are not specific to any microbial group and distorted some of the depth patterns. To assess the microbial community structure, the following grouping of PLFAs was used (Willers et al., 2015): Gram positive bacteria (Gram+; iC14:0, aC14:0, iC15:0, aC15:0, iC16:0, aC16:0, iC17:0, and aC17:0), Gram negative bacteria (Gram−; C16:1ω5c, C16:1ω7c, C16:1ω9c, C18:1ω5c, C18:1ω11c, cyC17:0, and cyC19:0), fungi (C18:2ω6c), and Actinobacteria (10MeC16:0 and 10MeC18:0). Microorganisms can also adapt their membrane composition in response to various stress factors, confounding physiological adaptations and changes in community structure (Wixon and Balser, 2013). The ratio of anteiso to iso branched PLFAs ((aC14:0 + aC15:0 + aC16:0 + aC17:0) (iC14:0 + iC15:0 + iC16:0 + iC17:0)) can partly capture the adaptations of the membrane composition within Gram+ bacteria in response to stress rather than changes in the community composition (Hall et al., 2010; Wixon and Balser, 2013). The difference in PLFAs between the warmed and control plots was calculated as the difference between the warmed and control treatments normalized to the control plot within each of the three replicate paired blocks.
2.2.3 Branched glycerol dialkyl glycerol tetraethers
Samples were extracted using Soxhlet extraction with 93:7 () dichloromethane : methanol, as described in Wiesenberg and Gocke (2017). The extract was then prepared for an analysis of core lipid brGDGTs, as described by Coffinet et al. (2014). Briefly, the samples were separated over a column of activated aluminum oxide in two fractions. Branched glycerol dialkyl glycerol tetraethers were separated using high-performance liquid chromatography equipped with an automatic injector, coupled with mass spectrometry with an atmospheric pressure chemical ionization source (HPLC-APCI-MS; Shimadzu LCMS-2020; Shimadzu Corporation). In total, two Hypersil Gold silica columns in tandem (150 mm × 2.1 mm and 1.9 µm; Thermo Finnigan LLC), thermally controlled at 40 ∘C, were used for brGDGT analysis, as described in detail in Huguet et al. (2019). Semi-quantification of brGDGTs was performed by comparing the integrated signal of the respective compound with the signal of a C46 synthesized internal standard, assuming their response factors to be identical (Huguet et al., 2013).
Roman numerals correspond to the different brGDGT structures presented in Fig. A1, where compounds denoted by an accent after the Roman numerals are 6-methyl isomers. The relative abundances of individual brGDGTs (weight percentage) were calculated by dividing individual compounds by the sum of all brGDGTs. The difference in brGDGTs between warmed and control plots was calculated as being the difference between the warmed and control treatments normalized to the control plot within each of the three replicate paired blocks.
2.3 Data analysis
For all statistical analyses, R 3.6.3 (R Core Team, 2020) was used. We assessed normality, homoscedasticity, and model fit using residual plots, Shapiro–Wilk test, Levene test, and outlier tests. We log transformed the data if we observed a poor fit. Correlation and multiple regression analyses were conducted to test the relation between MBC, PLFA, brGDGT, and carbon concentrations. To test how warming affected microbial abundance and community, mixed effect models were built using the gls and lme functions in the nlme package. We used restricted maximum likelihood with the random effect block (n=3) and the fixed effects treatment, depth, and their interaction. We used Akaike's information criterion (AIC) to test whether variance structures improved model fit. If we found significant treatment–depth interaction or, upon visual inspection of the plots, the model was run on a subset of depth increments. This is indicated by the specification of depth intervals included in the analysis. We reported results as significant at a level of p <0.05.
3.1 Bulk parameters
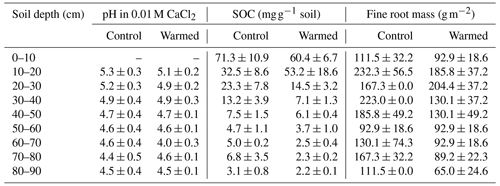
The pH was lower with depth (LME; p = 0.02; F = 3.04), with values ranging from 5.3 ± 0.3 at 10–20 cm depth and 4.5 ± 0.4 at 80–90 cm depth in control plots (Table 1). The pH was not different between control and warmed plots (LME; p = 0.84; F = 0.04). Soil organic carbon concentrations were also lower with depth (LME; p <0.01; F = 55.92), with values ranging from 71.3 ± 10.9 mg g−1 soil at 0–10 cm to 3.1 ± 0.8 mg g−1 soil at 80–90 cm depth in control plots (Table 1). Soil organic carbon concentrations were lower in warmed plots compared to control plots below 20 cm (LME; p <0.01; F = 15.17) in the soil cores analyzed here and as described in more detail on replicated cores in Soong et al. (2021). Fine root concentrations did not change with depth but were lower in warmed plots compared to control plots, as described in Ofiti et al. (2021; Table 1).
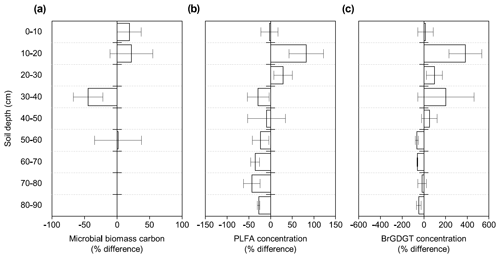
Figure 1Changes in microbial abundance between warmed and control plots based on (a) microbial biomass carbon using chloroform fumigation extraction (Soong et al., 2021), (b) PLFA concentrations, and (c) brGDGT concentrations after 4.5 years of +4 ∘C soil warming. Values are shown as [(warmed–control)/control × 100] (mean ± SE; n=3). The scale is adjusted to the respective values.
3.2 Microbial biomass carbon
Microbial biomass carbon (MBC) concentrations declined with increasing depth, independent of treatment (LME; p <0.01; F = 8.21). In the top 10 cm, concentrations were 727.6 ± 137.9 and 188.7 ± 98.2 ug C per gram of soil at 50–60 cm in control plots (Table B2). Microbial biomass carbon concentrations were below the detection limit for the depth increment 80–90 cm. Warming did not affect MBC (LME; p = 0.59; F = 0.31; Fig. 1). Concentrations of MBC and carbon concentrations were well correlated (R = 0.79; Fig. A2).
3.3 Phospholipid fatty acids
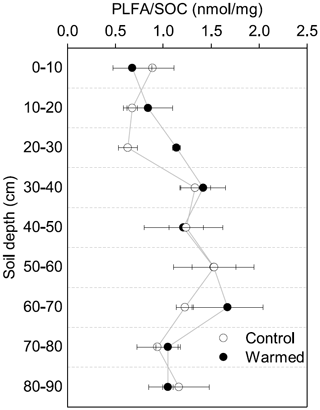
Similar to the MBC, PLFA concentrations declined with depth, independent of treatment (LME; p <0.01; F = 41.95). Concentrations were 62.4 ± 16.6 nmol g−1 soil in the top 10 cm of the mineral soil and 3.1 ± 0.2 nmol g−1 soil at 80–90 cm in control plots (Table B2). Concentrations of PLFAs and carbon and concentrations of PLFAs and MBC were well correlated (R = 0.94 and R = 0.82, respectively; Fig. A3a and b). The proportion of PLFAs to SOC increased with soil depth (p <0.01; Fig. 2).
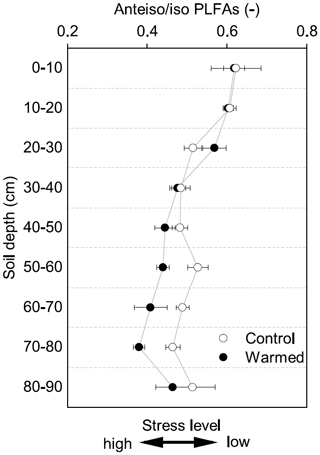
Figure 3The ratio of anteiso to iso PLFAs in control and warmed plots after 4.5 years of +4 ∘C soil warming (mean ± SE; n=3). This ratio has been interpreted as being an indicator of stress on Gram+ bacteria, with lower values indicating a higher stress level.
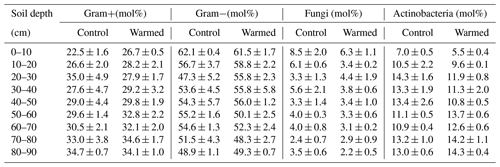
Gram− bacteria and fungi were relatively more abundant in the topsoil compared to subsoil (LME; where, respectively: p = 0.01, F = 3.06; p = 0.02, F = 2.68). In control plots, Gram− bacteria accounted for 62.1 ± 0.4 mol% in the top 10 cm and 48.9 ± 1.1 mol% at 80–90 cm and fungi for 8.5 ± 2.0 mol% and 3.5 ± 0.6 mol%, respectively (Table B3). Gram+ bacteria and Actinobacteria, on the other hand, were relatively more abundant in subsoil (LME; where, respectively: p <0.01, F = 3.43; p <0.01, F = 7.59). In control plots, Gram+ bacteria accounted for 22.5 ± 1.6 mol% in the top 10 cm and 34.7 ± 0.7 mol% at 80–90 cm and Actinobacteria for 7.0 ± 0.5 mol% and 13.0 ± 0.6 mol%, respectively (Table B3). The ratio of anteiso to iso PLFAs declined with depth (LME; p <0.01; F = 10.51; Fig. 3).
The concentrations of PLFAs were lower in warmed plots compared to control plots (LME; p = 0.045; F = 4.33; Fig. 1b). Below 30 cm, the concentrations of PLFAs were significantly lower by 28 % in warmed plots compared to control plots (LME30−90 cm; p <0.01; F = 9.69). The proportions of total PLFA concentrations to SOC were not affected by warming (LME; p = 0.51; F = 0.44; Fig. 2).
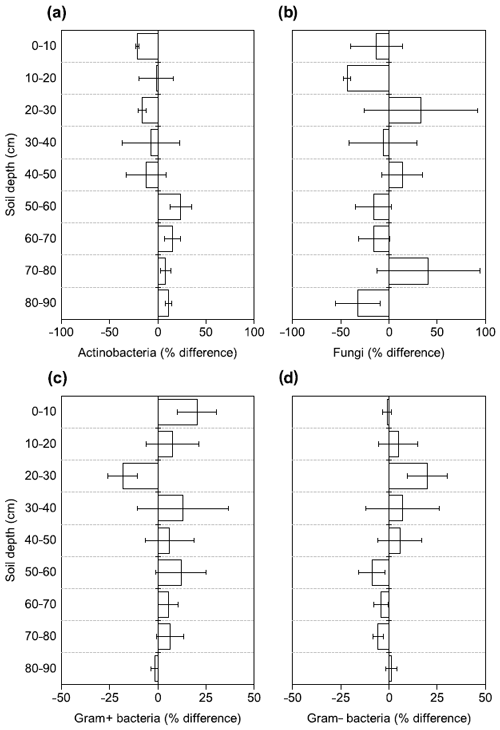
Figure 4Changes in the relative abundance of (a) Actinobacteria, (b) fungi, (c) Gram+ bacteria, and (d) Gram− bacteria based on PLFA biomarkers after 4.5 years of +4 ∘C soil warming. Values are shown as [(warmed–control)/control × 100] (mean ± SE; n=3).
There were no differences between warmed and control plots in the relative abundance of Gram+, Gram−, Actinobacteria and fungi over the whole-soil profile (LME; p = 0.34, F = 0.95; p = 0.72, F = 0.13; p = 0.58, F = 0.32 and p = 0.19, F = 1.76, respectively, Fig. 4). Below 50 cm, the relative abundance of Actinobacteria was higher in warmed plots compared to control plots (LME50−90 cm, p <0.01, F = 11.58; Fig. 4a). The ratio of anteiso to iso PLFAs was lower in warmed plots compared to control plots (LME, p = 0.01, F = 7.44; Fig. 3).
3.4 Branched glycerol dialkyl glycerol tetraethers
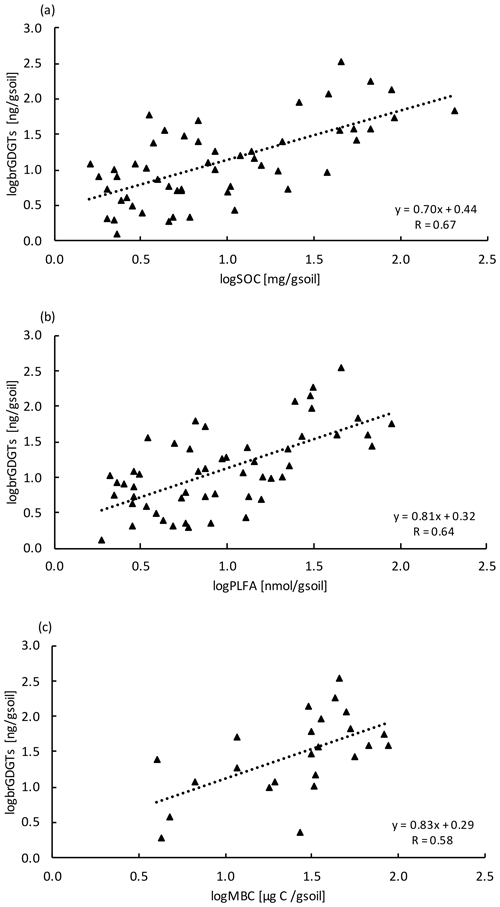
Concentrations of brGDGTs declined with depth (LME; p <0.01; F = 9.99). In the top 10 cm of control plots, concentrations were 88.7 ± 48.1 ng per gram of soil and, at 80–90 cm depth, 19.7 ± 8.2 ng per gram of soil (Table B2). There was only a moderate correlation between brGDGT concentrations and concentrations of SOC, PLFAs, or MBC (R = 0.67, R = 0.64, and R = 0.58, respectively; Fig. A4a–c).
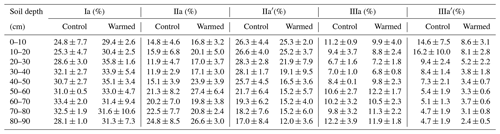
The relative abundance of individual brGDGTs compared to the total was dominated by compounds Ia, IIa, IIa′, IIIa, and IIIa′ (Table B4). Compounds IIa′ and IIIa′ were less abundant with depth (LME; p = 0.03, F = 2.46 and p <0.01, and F = 3.84, respectively; Table B4).
There was no effect of warming on the concentrations of brGDGTs over the whole-soil profile (LME; p = 0.28; F = 1.22; Fig. 1c). Below 50 cm, the concentrations of brGDGTs were lower in warmed plots compared to control plots (LME50−90 cm; p <0.01; F = 12.13). There was no effect of warming on the relative abundance of any individual brGDGT (LME; p >0.05 for all; Table B4).
4.1 Depth trend in microbial abundance and community structure
Both MBC and PLFA microbial abundance proxies indicated lower microbial concentrations in subsoils compared to topsoils, independent of treatment (Table B2). This depth trend strongly correlates with SOC concentrations (Fig. A2 and A3a), supporting the consensus that the concentration of SOC is an important controlling factor on microbial abundance in soils (Fierer, 2017; Soong et al., 2020b). Similarly, carbon availability is an important factor structuring the microbial community composition in soil depth profiles, with oligotrophic bacteria favored when nutrients become limiting (Brewer et al., 2019; Fierer et al., 2003b). Increasing relative abundance of Gram+ and decreasing abundance of Gram− bacteria with depth (Table B3) indicates a decrease in SOC availability (Fanin et al., 2019). Gram− bacteria are more dependent on labile plant-derived carbon sources, whereas Gram+ bacteria are able to grow on more complex carbon sources (Fanin et al., 2019). The observed relative increase in Actinobacteria with depth (Table B3) is consistent with previous findings from a grassland site (Fierer et al., 2003b). It has been shown that certain Actinobacteria are able to degrade complex carbon sources and grow mycelia-like structures, which might allow this microbial group to thrive in the subsoil (Goodfellow and Williams, 1983). Fungal abundance decreased with depth (Table B3), likely because fresh plant-derived carbon, which supports the growth of saprotrophic fungi, is more available in the upper soil horizons (Griffiths et al., 1999; Lindahl et al., 2007). Finally, the ratio of anteiso to iso fatty acids has been used as a stress indicator in Gram+ bacteria (Wixon and Balser, 2013), where a lower ratio indicates that the membrane fluidity is higher, which is an adaptation of microorganisms to maintain growth (Hall et al., 2010). The lower ratio in subsoils (Fig. 3) might be related to the low-carbon environment in subsoils. However, other factors such as pH, soil moisture, or temperature also shape the microbial community (Drenovsky et al., 2004; Fierer, 2017). For example, the pH at our site is lower in the subsoil compared to topsoil, which most likely will stimulate adaptations of the cell membranes.
The concentrations of brGDGTs also declined with depth (Table B2), but the correlation with carbon concentrations (Fig. A4a) and the other microbial biomass proxies (Fig. A4b,c) was only moderate (R <0.67). The moderate correlation is likely because brGDGTs are a time-integrated signal of accumulated necromass with a turnover time of several decades (Weijers et al., 2010). We did observe significant depth trends for two brGDGTs. However, it is unclear whether these trends reflect changes in the community structure or physiological adaptations of single microorganisms with depth as the source organisms are not yet fully identified (Sinninghe Damsté et al., 2014).
Overall, carbon concentrations seem to be a strong control on the microbial proxies MBC and PLFAs, whereas the brGDGT distribution is known to be strongly influenced by mean annual air temperature and pH but may also be affected by other factors like vegetation (Ayari et al., 2013), seasonality (Huguet et al., 2013), or soil moisture (Menges et al., 2014).
4.2 Less microbial biomass with warming in subsoil, but not topsoil
Our results show a distinctly different response of topsoil and subsoil microbial abundance to warming in PLFA concentrations (Fig. 1b). There was no significant effect of warming on the microbial abundance in the topsoil, which confirms previous observations from forest topsoils (Chen et al., 2015). Lower PLFA concentrations in warmed subsoils compared to control subsoils contradict our first hypothesis that the increased respiration observed at all depths (Hicks Pries et al., 2017; Soong et al., 2021) is at least partially reflected by an increase in microbial biomass. Lower concentrations of brGDGTs in warmed subsoils compared to control subsoils are in line with the observation in PLFAs, even though only at depths from 50 to 90 cm (Fig. 1c), and with our second hypothesis that microbial necromass would be less abundant in warmed soils. Due to the higher persistence of brGDGTs in soils, there might be a lag in response as compared to PLFAs (Weijers et al., 2010). Nevertheless, accelerated decomposition at soil depths below 50 cm, as observed by Ofiti et al. (2021), likely combined with lower microbial biomass inputs cause a decrease in concentrations of brGDGTs. Microbial biomass carbon, on the other hand, was not affected by warming at any depth (Fig. 1a). We hypothesize that the bulk measurement of MBC might be less powerful at capturing treatment effects compared to the targeted molecular analysis of PLFAs and brGDGTs.
The limitation of available carbon has been suggested as a potential driver of decreasing microbial abundance in long-term warming experiments (Kirschbaum, 2004; Walker et al., 2018) and has been reported in surface soils of field warming experiments (Melillo et al., 2017; Pold et al., 2017). Microbial abundance closely tracked carbon concentrations with depth and as they changed with warming. The ratio of total PLFAs to SOC did not change with warming (Fig. 2), despite 28 % lower PLFA concentrations in the warmed subsoil (Fig. 1b). Soil organic carbon inputs to subsoils, in the form of dissolved organic carbon (DOC) cascading down from the topsoil, are often strongly processed (Roth et al., 2019), and fine root litter inputs decline with depth (Leppälammi-Kujansuu et al., 2013). Declining SOC inputs with depth may limit microbially available carbon with warming in subsoils as compared to topsoils. We observed lower concentrations of particulate organic matter (Soong et al., 2021) and fine root mass in the warmed subsoils (Table 1; Ofiti et al., 2021). After 3.5 years of warming, unprotected SOC (i.e., not mineral associated) had been depleted due to warming in the subsoil (Soong et al., 2021), indicating a loss of easily accessible organic matter for subsoil microorganisms. These results suggest that, in this forest, carbon limitation might be one cause for the lower abundance of microorganisms in subsoil after 4.5 years of +4 ∘C warming.
In addition to SOC availability, temperature (Bradford, 2013) and changes in soil moisture associated with warming (Manzoni et al., 2012) likely affect microbial abundance. We did observe a similar respiration response at all depths (Hicks-Pries et al. 2017), indicating that the microbial response might be similar despite the difference in warming magnitude between the top 20 cm and the lower horizons. Nevertheless, temperature controls the reaction rates of microbial enzymes, which, in turn, can affect microbial abundance (Allison et al., 2010), and incubation experiments show strong effects of temperature magnitude on soil respiration, including in subsoil (Yan et al., 2017). Thus, the higher magnitude of warming below 20 cm in our experiment might be partially responsible for the observed difference in the microbial response between the top- and subsoil. Furthermore, warming of the soil caused a significant decrease in the volumetric water content by 2–5 percentage points throughout the soil profile in the warmed plots in our experiment (Soong et al., 2021). Drying is expected to coincide with rising temperatures in Mediterranean climate regions under future climate scenarios (Polade et al., 2017). In mesic soils, drying generally has a negative effect on microbial abundance and activity (Manzoni et al., 2012).
To our knowledge, only Zhang et al. (2015) reported microbial responses to in situ subsoil warming. In their experiment, warming induced an increase in microbial biomass down to 50 cm, and the authors proposed that this was due to increased SOC availability due to stimulated plant growth, a reduction in anaerobic conditions, and better thermal growth conditions for microorganisms (Zhang et al., 2015). The uniform increase in microbial biomass and the proposed underlying processes could not be observed in our experiment, perhaps because the warming treatment was different. Zhang et al. (2015) created a gradient of warming from +1.4 ∘C at 10 cm depth to only +0.1 ∘C at 50 cm depth, barely warming the subsoil. Warming throughout the profile may offer a treatment closer to predictions of whole-soil warming by climate change (Soong et al., 2020a). Furthermore, plant dynamics and environmental factors, such as temperature and soil hydrology, are very different between their grassland site on the Tibetan plateau and our forest site in a Mediterranean climate. Microorganisms at their site are much more limited by temperature (mean annual temperature, MAT – −3.8 ∘C) and affected by anoxic conditions (Zhang et al., 2015). We propose that, similar to topsoils, in subsoils the ecosystem type and experimental setup strongly affect the observed response of microbial abundance to warming (Chen et al., 2015).
4.3 Small effect on microbial community composition in subsoils
Whereas 4.5 years of +4 ∘C warming decreased subsoil microbial abundance by 28 %, there were only minor effects on the microbial community structure as assessed by organic geochemical methods. Only Actinobacteria increased in relative abundance with warming in the lower subsoil (below 50 cm; Fig. 4a). We also observed a decrease in the ratio of anteiso to iso fatty acids with warming (Fig. 3). Changes in this ratio have been attributed to physiological adaptations of Gram+ bacteria to warming-induced stress rather than shifts in the community structure (Wixon and Balser, 2013). The above observations are in line with our third hypothesis that warming would cause depth-dependent changes in the microbial community.
So far, laboratory incubations tentatively identified Actinobacteria as one of the microbial groups that can be stimulated by warming (Oliverio et al., 2017). Our field observation of higher relative abundance of Actinobacteria in the lower subsoils confirms the findings from these laboratory incubations and shows that warming affected Actinobacteria less negatively relative to the other microbial groups. Furthermore, our results are also consistent with observations by Frey et al. (2008). They reported increased relative abundance of Actinobacteria in a long-term warming experiment in topsoils after more than 12 years of warming, where lower SOC availability was a main driver of the microbial response (Frey et al., 2008). Characteristics of Actinobacteria indicate their ability to degrade complex polymers in soils (Goodfellow and Williams, 1983), and long-term experimental warming caused an increase in the carbohydrate degrading genes of Actinobacteria (Pold et al., 2016). Finally, Actinobacteria seem to be more resistant to drought than other microbial taxa (Naylor et al., 2017). It remains uncertain whether this is due to their ability for sporulation or other mechanisms (Naylor et al., 2017), but the resistance to decreasing soil moisture could further contribute to the relative increase in abundance in our warmed subsoils. Thus, Actinobacteria might be relatively more abundant due to a combination of resilience to warming and adaption to lower SOC concentrations and reduced soil moisture.
Although warming did not alter the relative abundance of Gram+ bacteria, it did significantly change the fatty acid composition of microbial membranes, expressed as the ratio of anteiso to iso branched phospholipid fatty acids in the subsoil (Fig. 3). The ratio of anteiso to iso fatty acids likely reflects an acclimation to changing temperatures in Gram+ bacteria, which might give Gram+ bacteria a competitive advantage because they can adapt to changing environmental conditions (Hall et al., 2010). Nonetheless, the synthesis of new lipids is energy intensive and might contribute to the lower efficiency at which carbon is incorporated to microbial biomass (Hall et al., 2010). Overall, multiple years of soil warming caused a shift in the microbial community at Blodgett Forest towards a higher relative abundance of organisms able to utilize complex organic compounds and physiologically adapt to higher temperatures. Thus, there could be continued enhancement of decomposition, including previously stable SOC, in warmed subsoils.
The shifts in the microbial community and physiology observed by PLFAs were not reflected in the relative abundance of individual brGDGTs (Table B4). Likewise, at a decadal geothermal temperature gradient, the brGDGT fingerprint only changed at a high warming magnitude where soil temperatures were > +14 ∘C above ambient and when the microbial community was also significantly altered (De Jonge et al., 2019). In our experiment, lower concentrations of brGDGTs in the subsoil (Fig. 1c) did not affect the relative abundance of individual brGDGTs. This indicates that, after 4.5 years of warming, there was no shift in the microbial community or physiology, which produced detectable shifts in brGDGTs, nor preferential decomposition of certain core brGDGTs.
Further research is needed to assess how microbial abundance and community composition will evolve with long-term subsoil warming. A long-term study at Harvard Forest indicated that the microbial response to warming, at least in topsoils, progressed in cycles of change in the microbial community and decomposition rates (Melillo et al., 2017). It is not known whether similar impacts will occur in subsoils. Furthermore, seasonality can affect microbial biomass and community structure (Contosta et al., 2015) and only having samples from one time point is a limitation in our study. The fact that subsoil microorganisms are differently affected by warming compared to topsoils is valuable information needed to predict the long-term response of ecosystems to climate change, even if these effects diminish over the long term or are seasonally varying.
Microbial biomass concentrations in warmed subsoil (below 30 cm) were 28 % lower compared to the control after 4.5 years of +4 ∘C warming, whereas they were not significantly affected in topsoils, consistent with changes in SOC stock. We showed for the first time that subsoil and topsoil microbial communities respond differently to whole-soil warming. In the lower subsoil (below 50 cm), Actinobacteria were relatively more abundant, whereas Gram+ bacteria revealed physiological adaptations of the cell membrane in response to warming-related stress. Changes in the amount and composition of subsoil SOC, along with the direct effects of warming, are likely the main drivers of changes in the microbial community. Whether less biomass in subsoils will lead to reduced respiration in the future remains to be investigated. The observed shift in the subsoil microbial community composition favored Actinobacteria, which are able to metabolize more complex carbon sources, suggesting that soil warming may lead to microbial responses that disproportionately enhance degradation of previously stable subsoil carbon.
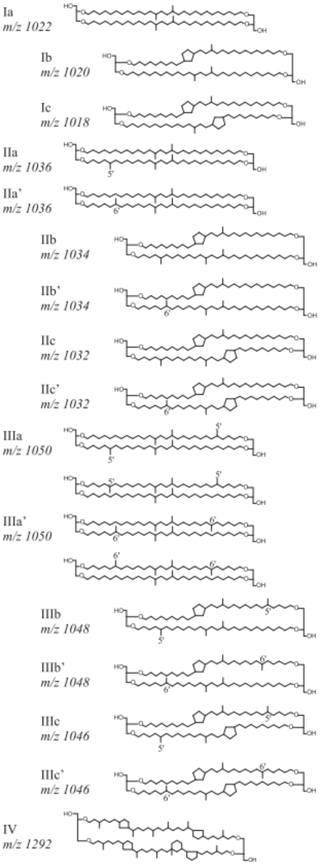
Figure A1Chemical structures and molecular masses of brGDGTs, as described in De Jonge et al. (2014).
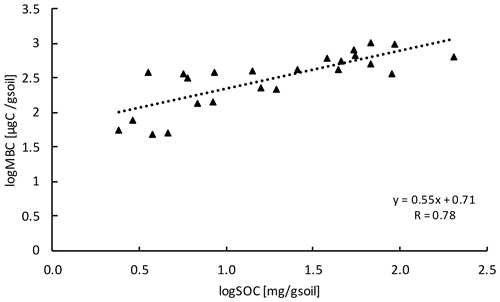
Figure A2Relationship between MBC and SOC at all depth increments (n=24). Values were log transformed to meet normality.
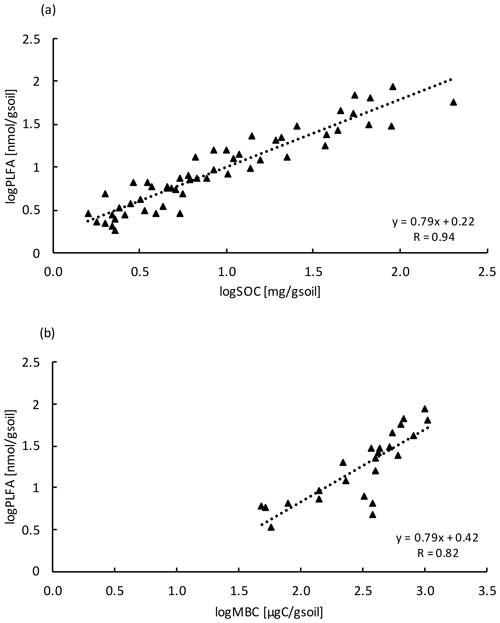
Figure A3Relationship between (a) PLFA with SOC (n=54) and (b) PLFA with MBC (n=24) at all depth increments. Values were log transformed to meet normality.
Table B2Concentrations of MBC using chloroform fumigation extraction (Soong et al., 2021), PLFAs, and brGDGTs in control plots and warmed plots (mean ± SE; n=3).
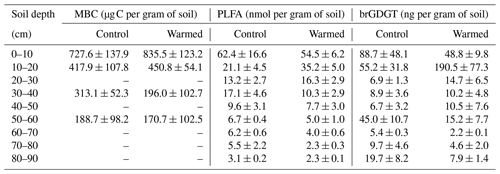
Table B3Relative abundance of microbial groups, identified using PLFAs in control plots and warmed plots (mean ± SE; n=3).
The data used in this study are available from the ESS-DIVE repository (https://doi.org/10.15485/1810163, Zosso et al., 2021).
MST designed and maintained the field warming experiment. MWIS conceived the DEEP C project. CUZ, NOEO, JLS, EFS, GLBW, MST, and MWIS participated in the field campaign. CUZ carried out biogeochemical analyses, supervised by GLBW and AH. All co-authors contributed to the data interpretation and to the paper, which was written by CUZ.
The authors declare that they have no conflict of interest.
Publisher's note: Copernicus Publications remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
We thank Cristina Castanha (LBL), for the field assistance, and Tatjana Kraut (UZH), for the laboratory assistance. Sylvie Derenne (Sorbonne Université) and Marijn van de Broek (University of Zurich) contributed to the advancement of our work with valuable comments and discussions. The University Research Priority Program Global Change and Biodiversity (URPP-GCB) at the University of Zurich offered opportunities for helpful discussions and exchange.
This research has been supported by the Swiss National Science Foundation (SNF; project no. 200021_172744), the U.S. Department of Energy Office of Science, the Office of Biological and Environmental Research Terrestrial Ecosystem Science Program (grant no. DE-SC-0001234), and the SATW Germaine de Staël (project no. 2019-08). Emily Solly acknowledges funding from the Swiss National Science Foundation (Ambizione; grant no. PZ00P2_180030).
This paper was edited by Ashish Malik and reviewed by Grace Pold and one anonymous referee.
Allison, S. D., Wallenstein, M. D., and Bradford, M. A.: Soil-carbon response to warming dependent on microbial physiology, Nat. Geosci., 3, 336–340, https://doi.org/10.1038/ngeo846, 2010.
Ayari, A., Yang, H., Xie, S., and Wiesenberg, G. L. B.: Distribution of archaeal and bacterial tetraether membrane lipids in rhizosphere-root systems in soils and their implication for paleoclimate assessment, Geochem. J., 47, 337–347, https://doi.org/10.2343/geochemj.2.0249, 2013.
Bai, T., Tao, J., Li, Z., Shu, M., Yan, X., Wang, P., Ye, C., Guo, H., Wang, Y., and Hu, S.: Different microbial responses in top- and sub-soils to elevated temperature and substrate addition in a semiarid grassland on the Loess Plateau, Eur. J. Soil Sci., 70, 1025–1036, https://doi.org/10.1111/ejss.12800, 2019.
Bradford, M. A.: Thermal adaptation of decomposer communities in warming soils, Front. Microbiol., 4, 333, https://doi.org/10.3389/fmicb.2013.00333, 2013.
Bradford, M. A., Davies, C. A., Frey, S. D., Maddox, T. R., Melillo, J. M., Mohan, J. E., Reynolds, J. F., Treseder, K. K., and Wallenstein, M. D.: Thermal adaptation of soil microbial respiration to elevated temperature, Ecol. Lett., 11, 1316–1327, https://doi.org/10.1111/j.1461-0248.2008.01251.x, 2008.
Brewer, T. E., Aronson, E. L., Arogyaswamy, K., Billings, S. A., Botthoff, J. K., Campbell, A. N., Dove, N. C., Fairbanks, D., Gallery, R. E., Hart, S. C., Kaye, J., King, G., Logan, G., Lohse, K. A., Maltz, M. R., Mayorga, E., O'Neill, C., Owens, S. M., Packman, A., Pett-Ridge, J., Plante, A. F., Richter, D. D., Silver, W. L., Yang, W. H., and Fierer, N.: Ecological and genomic attributes of novel bacterial taxa that thrive in subsurface soil horizons, MBio, 10, e01318-19, https://doi.org/10.1128/mBio.01318-19, 2019.
Chen, J., Luo, Y., Xia, J., Jiang, L., Zhou, X., Lu, M., Liang, J., Shi, Z., Shelton, S., and Cao, J.: Stronger warming effects on microbial abundances in colder regions, Sci. Rep.-UK, 5, 1–10, https://doi.org/10.1038/srep18032, 2015.
Coffinet, S., Huguet, A., Williamson, D., Fosse, C., and Derenne, S.: Potential of GDGTs as a temperature proxy along an altitudinal transect at Mount Rungwe (Tanzania), Org. Geochem., 68, 82–89, https://doi.org/10.1016/j.orggeochem.2014.01.004, 2014.
Contosta, A. R., Frey, S. D., and Cooper, A. B.: Soil microbial communities vary as much over time as with chronic warming and nitrogen additions, Soil Biol. Biochem., 88, 19–24, https://doi.org/10.1016/j.soilbio.2015.04.013, 2015.
De Jonge, C., Hopmans, E. C., Zell, C. I., Kim, J.-H., Schouten, S., and Sinninghe Damsté, J. S.: Occurrence and abundance of 6-methyl branched glycerol dialkyl glycerol tetraethers in soils: Implications for palaeoclimate reconstruction, Geochim. Cosmochim. Ac., 141, 97–112, https://doi.org/10.1016/j.gca.2014.06.013, 2014.
De Jonge, C., Radujković, D., Sigurdsson, B. D., Weedon, J. T., Janssens, I., and Peterse, F.: Lipid biomarker temperature proxy responds to abrupt shift in the bacterial community composition in geothermally heated soils, Org. Geochem., 137, 103897, https://doi.org/10.1016/j.orggeochem.2019.07.006, 2019.
Drenovsky, R. E., Vo, D., Graham, K. J., and Scow, K. M.: Soil water content and organic carbon availability are major determinants of soil microbial community composition, Microb. Ecol., 48, 424–430, https://doi.org/10.1007/s00248-003-1063-2, 2004.
Fanin, N., Kardol, P., Farrell, M., Nilsson, M.-C., Gundale, M. J., and Wardle, D. A.: The ratio of Gram-positive to Gram-negative bacterial PLFA markers as an indicator of carbon availability in organic soils, Soil Biol. Biochem., 128, 111–114, https://doi.org/10.1016/j.soilbio.2018.10.010, 2019.
Fierer, N.: Embracing the unknown: Disentangling the complexities of the soil microbiome, Nat. Rev. Microbiol., 15, 579–590, https://doi.org/10.1038/nrmicro.2017.87, 2017.
Fierer, N., Allen, A. S., Schimel, J. P., and Holden, P. A.: Controls on microbial CO2 production: A comparison of surface and subsurface soil horizons, Glob. Change Biol., 9, 1322–1332, https://doi.org/10.1046/j.1365-2486.2003.00663.x, 2003a.
Fierer, N., Schimel, J. P., and Holden, P. A.: Variations in microbial community composition through two soil depth profiles, Soil Biol. Biochem., 35, 167–176, https://doi.org/10.1016/S0038-0717(02)00251-1, 2003b.
Frey, S. D., Drijber, R., Smith, H., and Melillo, J.: Microbial biomass, functional capacity, and community structure after 12 years of soil warming, Soil Biol. Biochem., 40, 2904–2907, https://doi.org/10.1016/j.soilbio.2008.07.020, 2008.
Frostegård, Å., Tunlid, A., and Bååth, E.: Microbial biomass measured as total lipid phosphate in soils of different organic content, J. Microbiol. Meth., 14, 151–163, https://doi.org/10.1016/0167-7012(91)90018-L, 1991.
Gocke, M. I., Huguet, A., Derenne, S., Kolb, S., Dippold, M. A., and Wiesenberg, G. L. B.: Disentangling interactions between microbial communities and roots in deep subsoil, Sci. Total Environ., 575, 135–145, https://doi.org/10.1016/j.scitotenv.2016.09.184, 2017.
Goodfellow, M. and Williams, S. T.: Ecology of Actinomycetes, Annu. Rev. Microbiol., 37, 189–216, https://doi.org/10.1146/annurev.mi.37.100183.001201, 1983.
Griffiths, B. S., Ritz, K., Ebblewhite, N., and Dobson, G.: Soil microbial community structure: Effects of substrate loading rates, Soil Biol. Biochem., 31, 145–153, https://doi.org/10.1016/s0038-0717(98)00117-5, 1999.
Hall, E. K., Singer, G. A., Kainz, M. J., and Lennon, J. T.: Evidence for a temperature acclimation mechanism in bacteria: An empirical test of a membrane-mediated trade-off, Funct. Ecol., 24, 898–908, https://doi.org/10.1111/j.1365-2435.2010.01707.x, 2010.
Hicks Pries, C. E., Castanha, C., Porras, R. C., and Torn, M. S.: The whole-soil carbon flux in response to warming, Science, 355, 1420–1423, https://doi.org/10.1126/science.aal1319, 2017.
Huguet, A., Fosse, C., Laggoun-Défarge, F., Delarue, F., and Derenne, S.: Effects of a short-term experimental microclimate warming on the abundance and distribution of branched GDGTs in a French peatland, Geochim. Cosmochim. Ac., 105, 294–315, https://doi.org/10.1016/j.gca.2012.11.037, 2013.
Huguet, A., Coffinet, S., Roussel, A., Gayraud, F., Anquetil, C., Bergonzini, L., Bonanomi, G., 836 Williamson, D., Majule, A., and Derenne, S.: Evaluation of 3-hydroxy fatty acids as a pH and temperature proxy in soils from temperate and tropical altitudinal gradients, Org. Geochem., 129, 1–13, https://doi.org/10.1016/j.orggeochem.2019.01.002, 2019.
IPCC: Climate Change 2013: The Physical Science Basis. Contribution of Working Group I to the Fifth Assessment Report of the Intergovernmental Panel on Climate Change, edited by: Stocker, T. F., Qin, D., Plattner, G.-K., Tignor, M., Allen, S. K., Boschung, J, Nauels, A., Xia, Y., Bex, V., and Midgley, P. M., Cambridge University Press, Cambridge, UK and New York, NY, USA, 2013.
Jansson, J. K. and Hofmockel, K. S.: Soil microbiomes and climate change, Nat. Rev. Microbiol., 18, 35–46, https://doi.org/10.1038/s41579-019-0265-7, 2020.
Jobbagy, E. G. and Jackson, R. B.: The vertical distribution of soil organic carbon and its relation to climate and vegetation, Ecol. Appl., 10, 423–436, https://doi.org/10.1890/1051-0761(2000)010[0423:TVDOSO]2.0.CO;2, 2000.
Kindler, R., Miltner, A., Thullner, M., Richnow, H. H., and Kästner, M.: Fate of bacterial biomass derived fatty acids in soil and their contribution to soil organic matter, Org. Geochem., 40, 29–37, https://doi.org/10.1016/j.orggeochem.2008.09.005, 2009.
Kirschbaum, M. U. F.: Soil respiration under prolonged soil warming: Are rate reductions caused by acclimation or substrate loss?, Glob. Change Biol., 10, 1870–1877, https://doi.org/10.1111/j.1365-2486.2004.00852.x, 2004.
Leppälammi-Kujansuu, J., Ostonen, I., Strömgren, M., Nilsson, L. O., Kleja, D. B., Sah, S. P., and Helmisaari, H. S.: Effects of long-term temperature and nutrient manipulation on Norway spruce fine roots and mycelia production, Plant Soil, 366, 287–303, https://doi.org/10.1007/s11104-012-1431-0, 2013.
Lin, W., Li, Y., Yang, Z., Giardina, C. P., Xie, J., Chen, S., Lin, C., Kuzyakov, Y., and Yang, Y.: Warming exerts greater impacts on subsoil than topsoil CO2 efflux in a subtropical forest, Agr. Forest. Meteorol., 263, 137–146, https://doi.org/10.1016/j.agrformet.2018.08.014, 2018.
Lindahl, B. D., Ihrmark, K., Boberg, J., Trumbore, S. E., Högberg, P., Stenlid, J., and Finlay, R. D.: Spatial separation of litter decomposition and mycorrhizal nitrogen uptake in a boreal forest, New Phytol., 173, 611–620, https://doi.org/10.1111/j.1469-8137.2006.01936.x, 2007.
Manzoni, S., Schimel, J. P., and Porporato, A.: Responses of soil microbial communities to water stress: Results from a meta-analysis, Ecology, 93, 930–938, https://doi.org/10.1890/11-0026.1, 2012.
Melillo, J. M., Frey, S. D., DeAngelis, K. M., Werner, W. J., Bernard, M. J., Bowles, F. P., Pold, G., Knorr, M. A., and Grandy, A. S.: Long-term pattern and magnitude of soil carbon feedback to the climate system in a warming world, Science, 358, 101–105, https://doi.org/10.1126/science.aan2874, 2017.
Menges, J., Huguet, C., Alcañiz, J. M., Fietz, S., Sachse, D., and Rosell-Melé, A.: Influence of water availability in the distributions of branched glycerol dialkyl glycerol tetraether in soils of the Iberian Peninsula, Biogeosciences, 11, 2571–2581, https://doi.org/10.5194/bg-11-2571-2014, 2014.
Naylor, D., DeGraaf, S., Purdom, E., and Coleman-Derr, D.: Drought and host selection influence bacterial community dynamics in the grass root microbiome, ISME J., 11, 2691–2704, https://doi.org/10.1038/ismej.2017.118, 2017.
Nottingham, A. T., Meir, P., Velasquez, E., and Turner, B. L.: Soil carbon loss by experimental warming in a tropical forest, Nature, 584, 234–237, https://doi.org/10.1038/s41586-020-2566-4, 2020.
Ofiti, N. O. E., Zosso, C. U., Soong, J. L., Solly, E. F., Margaret, S., and Schmidt, M. W. I.: Warming promotes loss of subsoil carbon through accelerated degradation of plant-derived organic matter, Soil Biol. Biochem., 156, 108185, https://doi.org/10.1016/j.soilbio.2021.108185, 2021.
Oliverio, A. M., Bradford, M. A., and Fierer, N.: Identifying the microbial taxa that consistently respond to soil warming across time and space, Glob. Change Biol., 23, 2117–2129, https://doi.org/10.1111/gcb.13557, 2017.
Polade, S. D., Gershunov, A., Cayan, D. R., Dettinger, M. D., and Pierce, D. W.: Precipitation in a warming world: Assessing projected hydro-climate changes in California and other Mediterranean climate regions, Sci. Rep.-UK, 7, 1–10, https://doi.org/10.1038/s41598-017-11285-y, 2017.
Pold, G., Billings, A. F., Blanchard, J. L., Burkhardt, D. B., Frey, S. D., Melillo, J. M., Schnabel, J., van Diepen, L. T. A., and DeAngelis, K. M.: Long-term warming alters carbohydrate degradation potential in temperate forest soils, Appl. Environ. Microbiol., 82, 6518–6530, https://doi.org/10.1128/AEM.02012-16, 2016.
Pold, G., Grandy, A. S., Melillo, J. M., and DeAngelis, K. M.: Changes in substrate availability drive carbon cycle response to chronic warming, Soil Biol. Biochem., 110, 68–78, https://doi.org/10.1016/j.soilbio.2017.03.002, 2017.
R Core Team: R: A language and environment for statistical computing, R Foundation for Statistical Computing, Vienna, Austria, available at: https://www.r-project.org/ (last access: 18 May 2021), 2020.
Ramsey, P. W., Rillig, M. C., Feris, K. P., Holben, W. E., and Gannon, J. E.: Choice of methods for soil microbial community analysis: PLFA maximizes power compared to CLPP and PCR-based approaches, Pedobiologia (Jena), 50, 275–280, https://doi.org/10.1016/j.pedobi.2006.03.003, 2006.
Roth, V.-N., Lange, M., Simon, C., Hertkorn, N., Bucher, S., Goodall, T., Griffiths, R. I., Mellado-Vázquez, P. G., Mommer, L., Oram, N. J., Weigelt, A., Dittmar, T., and Gleixner, G.: Persistence of dissolved organic matter explained by molecular changes during its passage through soil, Nat. Geosci., 12, 755–761, https://doi.org/10.1038/s41561-019-0417-4, 2019.
Rumpel, C. and Kögel-Knabner, I.: Deep soil organic matter-a key but poorly understood component of terrestrial C cycle, Plant Soil, 338, 143–158, https://doi.org/10.1007/s11104-010-0391-5, 2011.
Sinninghe Damsté, J. S., Rijpstra, W. I. C., Hopmans, E. C., Foesel, B. U., Wüst, P. K., Overmann, J., Tank, M., Bryant, D. A., Dunfield, P. F., Houghton, K., and Stott, M. B.: Ether- and ester-bound iso-diabolic acid and other lipids in members of Acidobacteria subdivision 4, Appl. Environ. Microbiol., 80, 5207–5218, https://doi.org/10.1128/AEM.01066-14, 2014.
Soong, J. L., Phillips, C. L., Ledna, C., Koven, C. D., and Torn, M. S.: CMIP5 Models Predict Rapid and Deep Soil Warming Over the 21st Century, J. Geophys. Res.-Biogeo., 125, 1–17, https://doi.org/10.1029/2019jg005266, 2020a.
Soong, J. L., Fuchslueger, L., Marañon-Jimenez, S., Torn, M. S., Janssens, I. A., Penuelas, J., and Richter, A.: Microbial carbon limitation: The need for integrating microorganisms into our understanding of ecosystem carbon cycling, Glob. Change Biol., 26, 1953–1961, https://doi.org/10.1111/gcb.14962, 2020b.
Soong, J. L., Castanha, C., Hicks Pries, C. E., Ofiti, N., Porras, R., Riley, W. J., Schmidt, M. W. I., and Torn, M. S.: Five years of whole-soil warming led to loss of subsoil carbon stocks and increased CO2 efflux, Science Advances, 7, 1–9, https://doi.org/10.1126/sciadv.abd1343, 2021.
Vance, E. D., Brookes, P. C., and Jenkinson, D. S.: An extraction Method for measuring soil microbial biomass C, Soil Biol. Biochem., 19, 703–707, https://doi.org/10.1016/0038-0717(87)90052-6, 1987.
Walker, T. W. N., Kaiser, C., Strasser, F., Herbold, C. W., Leblans, N. I. W., Woebken, D., Janssens, I. A., Sigurdsson, B. D., and Richter, A.: Microbial temperature sensitivity and biomass change explain soil carbon loss with warming, Nat. Clim. Change, 8, 885–889, https://doi.org/10.1038/s41558-018-0259-x, 2018.
Weijers, J. W. H., Wiesenberg, G. L. B., Bol, R., Hopmans, E. C., and Pancost, R. D.: Carbon isotopic composition of branched tetraether membrane lipids in soils suggest a rapid turnover and a heterotrophic life style of their source organism(s), Biogeosciences, 7, 2959–2973, https://doi.org/10.5194/bg-7-2959-2010, 2010.
Wiesenberg, G. L. B. and Gocke, M. I.: Analysis of Lipids and Polycyclic Aromatic Hydrocarbons as Indicators of Past and Present (Micro)Biological Activity, in: Hydrocarbon and lipid microbiology protocols, edited by: McGenity, T., Timmis, K., and Nogales, B., Springer, Berlin, Heidelberg, Germany, 61–91, https://doi.org/10.1007/8623_2015_157, 2017.
Willers, C., Jansen van Rensburg, P. J., and Claassens, S.: Phospholipid fatty acid profiling of microbial communities-a review of interpretations and recent applications, J. Appl. Microbiol., 119, 1207–1218, https://doi.org/10.1111/jam.12902, 2015.
Wixon, D. L. and Balser, T. C.: Toward conceptual clarity: PLFA in warmed soils, Soil Biol. Biochem., 57, 769–774, https://doi.org/10.1016/j.soilbio.2012.08.016, 2013.
Yan, D., Li, J., Pei, J., Cui, J., Nie, M., and Fang, C.: The temperature sensitivity of soil organic carbon decomposition is greater in subsoil than in topsoil during laboratory incubation, Sci. Rep.-UK, 7, 1–9, https://doi.org/10.1038/s41598-017-05293-1, 2017.
Zhang, B., Chen, S. Y., Zhang, J. F., He, X. Y., Liu, W. J., Zhao, Q., Zhao, L., and Tian, C. J.: Depth-related responses of soil microbial communities to experimental warming in an alpine meadow on the Qinghai-Tibet Plateau, Eur. J. Soil Sci., 66, 496–504, https://doi.org/10.1111/ejss.12240, 2015.
Zosso, C. U. and Wiesenberg, G. L. B.: Methylation procedures affect PLFA results more than selected extraction parameters, J. Microbiol. Meth., 182, 106164, https://doi.org/10.1016/j.mimet.2021.106164, 2021.
Zosso, C., Schmidt, M. W., Torn, M. S., Wiesenberg, G. L., Ofiti, N. O., Huguet, A., Soong, J. L., and Solly, E. F.: Blodgett Forest CA Warming Experiment phospholipid fatty acid and branched glycerol dialkyl glycerol tetraethers data 2018 from: “Whole-soil warming decreases abundance and modifies the community structure of microorganisms in the subsoil but not in surface soil”, Terrestrial Ecosystem Science at Berkeley Lab [data set], https://doi.org/10.15485/1810163, 2021.